Peptide hormone research models: a researcher’s guide
Peptide hormones are frequently treated as stable, well-characterized molecules in experimental design, but that assumption is one of the most consequential errors in biomedical research. Understanding what is a peptide hormone research model requires confronting a more complex reality: these signaling molecules are synthesized as inactive precursors, subjected to tissue-specific proteolytic cleavage, modified by post-translational processing, and subject to rapid degradation that can begin within seconds of sample collection. Each of these variables must be accounted for in a research model that produces reliable, interpretable data. This article provides a structured, technically grounded overview of the model types, methodological requirements, and experimental considerations that define rigorous peptide hormone research today.
Table of Contents
- Fundamentals of peptide hormone biology and experimental modeling
- Mass-spectrometry peptidomics as a core discovery model
- Receptor-level in vitro and in vivo models for functional peptide hormone study
- Computational and pharmacokinetic models advancing peptide hormone research
- In vivo receptor attribution models using conditional genetics
- Rethinking peptide hormone research models: integrating biology, technology, and experimental rigor
- Explore Vertex Labs’ solutions for advanced peptide research modeling
- Frequently asked questions
Key Takeaways
| Point | Details |
|---|---|
| Peptide processing complexity | Accurate models must account for precursor proteolysis, PTMs, and rapid degradation to reflect true peptide hormone biology. |
| Sample preparation criticality | Preventing peptide degradation during extraction is essential to avoid artifacts dominating mass spectrometry data. |
| Receptor-specific assays | Designing studies around specific receptor classes and signaling pathways ensures valid functional readouts. |
| Computational-experimental integration | In silico cleavage predictions require rigorous experimental validation to confirm peptide bioactivity. |
| Conditional genetic models | Targeted receptor knockouts clarify physiological effects in specific tissues, enhancing model precision. |
Fundamentals of peptide hormone biology and experimental modeling
What is a peptide hormone research model, at its core? It is a defined experimental system that accounts for the full biological lifecycle of a peptide hormone, from precursor synthesis to receptor engagement, within a controlled and reproducible framework. Before selecting any model, you must understand the biochemistry that makes these molecules difficult to study.
Peptide hormones are processed from larger precursors00063-5) through regulated proteolysis, with significant post-translational modifications (PTMs) that directly affect function and detection. Insulin, for example, is cleaved from proinsulin by prohormone convertases in the endoplasmic reticulum and Golgi apparatus. The resulting mature form bears no resemblance in mass or charge to its precursor. Detecting “insulin” without accounting for this processing means your assay may be measuring the wrong species entirely.
PTMs compound this problem substantially. Phosphorylation alters receptor binding affinity. Glycosylation affects circulating half-life and immunoreactivity. Amidation at the C-terminus, present in peptides like substance P and neuropeptide Y, is required for full biological activity. An experimental model that ignores these modifications will systematically underestimate or mischaracterize hormone potency.
Key considerations when establishing the biological foundation of your research model include:
- Precursor identity: Identify the prohormone and the specific convertases responsible for tissue-specific cleavage
- PTM profiling: Characterize modifications relevant to your peptide of interest before designing detection assays
- Degradation controls: Include protease inhibitor cocktails and temperature controls during sample collection
- Tissue specificity: Recognize that the same precursor can yield different mature peptides in different tissues
- Assay validation: Confirm that antibody-based or MS-based detection methods recognize the mature, modified form
Proper sample preparation best practices are not a procedural afterthought. They are a foundational element of the research model itself.
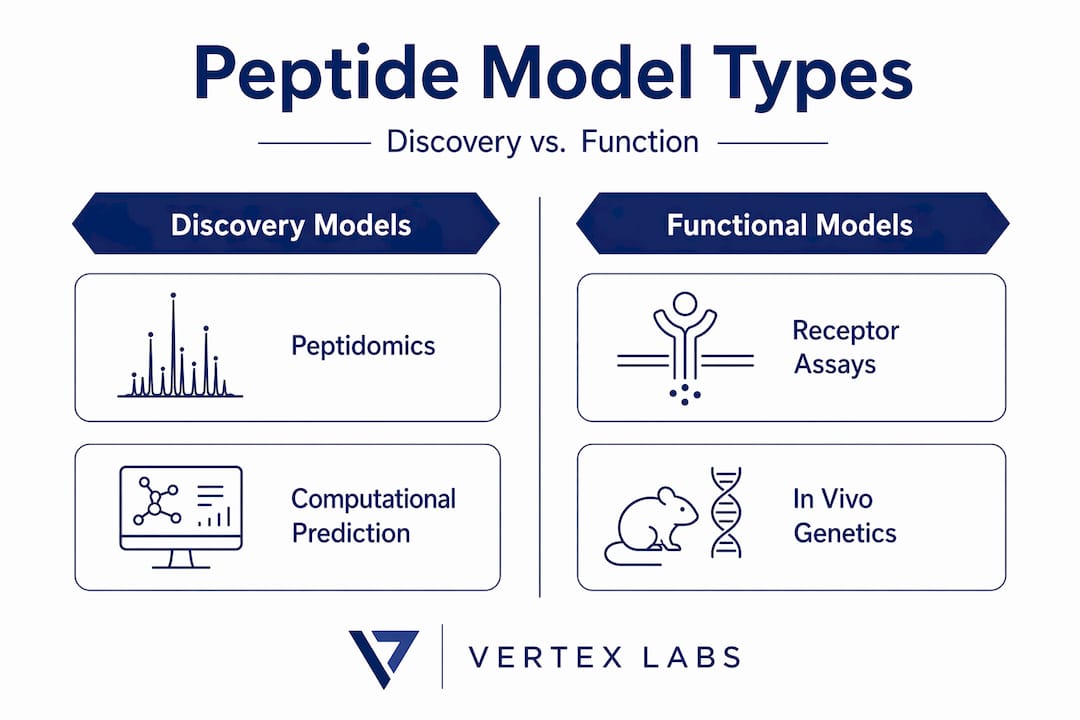
Mass-spectrometry peptidomics as a core discovery model
With biological fundamentals in place, we next examine how peptidomics experimentally captures peptide hormone profiles and the technical pitfalls researchers must avoid.
Mass spectrometry-based peptidomics has become the primary discovery tool for characterizing peptide hormone profiles in biological samples. It offers unbiased, high-throughput detection of peptides and their modifications across complex matrices including plasma, cerebrospinal fluid, and tissue homogenates. However, the method carries a critical and often underappreciated limitation.

More than 95% of peptides detected by MS00063-5) in standard tissue preparations are degradation fragments, not mature bioactive hormones. This is not a minor confound. It fundamentally alters the conclusions you can draw from peptidomics data unless active steps are taken to preserve the native peptide profile at the moment of sample collection.
Rapid heat inactivation using focused microwave irradiation is currently the most effective method for halting enzymatic degradation in tissue samples. The technique denatures proteases within milliseconds, preserving the endogenous peptide landscape. Without this step, the peptide profile you measure reflects post-mortem enzymatic activity rather than in vivo biology.
Bioinformatics pipelines and AI-assisted prediction tools add another layer of complexity. These platforms identify candidate bioactive peptides from large MS datasets by cross-referencing cleavage site databases and known prohormone sequences. They are powerful but require experimental validation before any candidate is treated as biologically relevant.
Pro Tip: When designing a peptidomics workflow, treat sample inactivation as a variable in your experimental design, not a standard operating procedure. Comparing inactivated versus non-inactivated samples from the same tissue can reveal how much of your detected peptide signal is artifactual, giving you a direct measure of model reliability.
Key methodological requirements for MS-based peptide hormone research models:
- Inactivation method: Implement focused microwave irradiation or boiling water immersion immediately post-dissection
- Inhibitor panels: Use broad-spectrum protease inhibitor cocktails as a secondary control
- Data filtering: Apply bioinformatics filters to distinguish mature hormones from degradation fragments
- Quantification standards: Use isotopically labeled internal standards for absolute quantification
- Replication: Run biological replicates to distinguish true peptide signals from technical noise
Receptor-level in vitro and in vivo models for functional peptide hormone study
Understanding how peptide hormones signal prepares us to examine the experimental models used to connect peptide presence with biological effects.
Detection of a peptide hormone in a sample tells you what is present. It does not tell you what it does. Functional characterization requires receptor-level models that link ligand exposure to defined signaling outcomes. The design of these models depends entirely on the receptor class your peptide engages.
Peptide hormones bind to three major receptor classes: G protein-coupled receptors (GPCRs), enzyme-coupled receptors such as receptor tyrosine kinases, and ligand-gated ion channel receptors. Each class activates distinct second messenger cascades, and each requires a different assay readout to accurately measure functional response.
A structured approach to receptor-level model selection:
- Identify the receptor class for your peptide of interest using existing literature and binding databases
- Select the appropriate assay readout: cAMP accumulation for Gs-coupled GPCRs, IP1 accumulation for Gq-coupled GPCRs, phosphorylation assays for receptor tyrosine kinases
- Choose a cell line with validated receptor expression or engineer stable expression in a null background to avoid confounding endogenous receptor activity
- Design concentration-response experiments across a physiologically relevant range to accurately determine EC50 values
- Include receptor antagonist controls to confirm that observed effects are receptor-mediated rather than off-target
- Transition to in vivo models only after in vitro receptor pharmacology is characterized, using conditional knockout animals to isolate receptor-specific contributions
Selecting the wrong assay for a given receptor class is among the most common sources of false negatives in peptide hormone studies. A peptide acting through a Gi-coupled receptor will suppress cAMP, not increase it. Running a standard cAMP stimulation assay will produce a null result and incorrectly classify an active compound as inactive.
Pro Tip: For GPCRs with known biased agonism, run parallel assays for both G protein and beta-arrestin pathways. Some peptide hormones preferentially engage one pathway over the other, and a single-readout assay will miss this pharmacological distinction entirely.
Computational and pharmacokinetic models advancing peptide hormone research
Beyond experimental and cellular models, computational and pharmacokinetic (PK) models provide predictive frameworks essential for validating peptide hormone candidates and understanding systemic behavior.
Computational cleavage prediction tools map potential mature peptide products from a given prohormone sequence by modeling known convertase recognition sites. A recent study demonstrated that computational drug discovery identified over 2,600 peptide fragments from PCSK1/3 cleavage of prohormone sequences, leading to the identification of a novel anti-obesity peptide. The computational step was essential for narrowing the candidate space. But it was in vivo validation that confirmed biological relevance.
Mechanistic PK models address a different but equally important question: once a peptide hormone enters systemic circulation, how does it distribute and clear? Mechanistic PK models use intravenous dosing data alongside in vitro clearance assays to estimate volume of distribution, clearance rate, and half-life. These parameters are then used to design dosing regimens in preclinical studies.
| Model type | Primary application | Key inputs | Common challenges |
|---|---|---|---|
| Computational cleavage | Candidate peptide identification | Prohormone sequence, convertase specificity | Requires experimental validation |
| Mechanistic PK | Systemic exposure prediction | IV dosing data, in vitro clearance | Parameter uncertainty, overprediction |
| Receptor binding simulation | Affinity and selectivity estimation | Structural data, docking algorithms | Conformational flexibility of peptides |
| Physiologically based PK | Tissue distribution modeling | Organ volumes, blood flow rates | Data-intensive, species translation gaps |
Iterative model refinement is standard practice, not a sign of model failure. Deviations between predicted and observed PK behavior direct you toward uncharacterized clearance mechanisms or tissue binding effects that require additional experimental characterization.
Pro Tip: When your mechanistic PK model overpredicts plasma exposure, the first variable to interrogate is renal filtration. Peptides below approximately 30 kDa are subject to glomerular filtration, and many PK models underestimate this clearance route without explicit renal compartment modeling.
Key applications of computational and PK modeling in peptide hormone research:
- Narrowing candidate peptide lists before costly in vivo studies
- Predicting tissue-specific exposure to guide receptor model selection
- Informing dosing schedules for conditional knockout experiments
- Supporting translational extrapolation from rodent to primate systems
In vivo receptor attribution models using conditional genetics
Having established modeling from biochemical to systemic levels, we conclude with advanced genetic tools for in vivo receptor attribution that increase model precision.
Conditional receptor knockout models represent the current standard for attributing specific physiological effects to defined receptor populations in intact organisms. Unlike whole-body knockouts, which eliminate receptor expression globally and introduce compensatory adaptations, conditional knockouts restrict gene deletion to specific cell types or tissues using Cre-lox recombination systems.
Conditional CNS GLP-1 receptor knockout mice provided direct evidence that central nervous system GLP-1 receptor signaling is required for the full weight-loss effect of a GIPR-antibody/GLP-1 peptide conjugate in obese mice. This finding could not have been obtained through pharmacological blockade alone, because GLP-1 receptor antagonists act at all receptor populations simultaneously, obscuring tissue-specific contributions.
Key design principles for conditional receptor attribution studies:
- Cre driver selection: Choose a Cre line with validated, tissue-restricted expression and confirm recombination efficiency before initiating studies
- Dosing protocol design: Use time-limited peptide exposure to capture acute receptor-mediated effects before compensatory mechanisms engage
- Quantitative phenotyping: Define primary endpoints (body weight, blood glucose, hormone levels) with sufficient statistical power before the study begins
- Littermate controls: Use Cre-negative littermates expressing the floxed allele as controls to account for Cre-associated toxicity
- Pathway confirmation: Validate receptor deletion at the protein level in target tissues using immunohistochemistry or flow cytometry
These models require substantial investment in experimental design and genetic validation. But they deliver a level of mechanistic clarity that no pharmacological approach can match.
Rethinking peptide hormone research models: integrating biology, technology, and experimental rigor
The field of peptide hormone research has a persistent problem with oversimplification. Researchers frequently select models based on convenience rather than biological fidelity. A radioimmunoassay designed for the intact hormone is applied to a sample that was processed at room temperature for 45 minutes. A cAMP assay is used to characterize a peptide acting through a Gi-coupled receptor. A whole-body knockout is used to attribute an effect that is actually tissue-specific. Each of these decisions produces data that is internally consistent but biologically misleading.
Most models risk oversimplification00063-5) by ignoring extensive degradation and processing complexity. The consequence is not just imprecise data. It is conclusions that cannot be replicated, candidate peptides that fail in subsequent studies, and mechanistic hypotheses that collapse under further scrutiny.
We believe the most productive reframing is to treat sample handling as part of the research model, not a preliminary step before the model begins. The moment you collect tissue or blood, enzymatic activity is shaping your data. Every minute at room temperature, every freeze-thaw cycle, every pH shift is a variable in your experiment. Researchers who internalize this produce fundamentally more reliable peptide hormone data.
Computational predictions deserve the same critical scrutiny. A list of 2,600 candidate peptides from a cleavage prediction algorithm is a starting point, not a finding. The value of computational modeling lies in its ability to direct experimental resources efficiently, not in replacing them. Tethering every computational prediction to pharmacological receptor assays and in vivo functional validation is not optional rigor. It is the minimum standard for credible peptide hormone research.
Conditional receptor knockouts provide superior insights over bulk pharmacologic approaches precisely because peptide hormone biology is inherently multi-tissue and context-dependent. The same peptide can have opposing effects in different tissues depending on receptor subtype expression and local signaling environment. Models that ignore this complexity will consistently produce results that fail to translate.
Integrating biological insight with methodologically appropriate tools and rigorous experimental controls is not an aspirational standard. It is the operational requirement for peptide hormone research that advances the field rather than adding noise to it.
Explore Vertex Labs’ solutions for advanced peptide research modeling
Designing a reliable peptide hormone research model depends on the quality of the compounds and protocols at the foundation of your workflow. Degradation artifacts, inconsistent purity, and undocumented batch variation are among the most common sources of irreproducible data in peptide studies.
At Vertex Peptides Lab, we provide laboratory-grade synthetic peptides verified at greater than 99% purity through third-party testing, with Certificates of Analysis available for every batch. Our catalog includes research compounds such as IGF-1 LR3, Ipamorelin, and TB-500, all synthesized under controlled conditions and supported by full batch documentation. For researchers building or refining peptide research workflows, our platform offers the compound integrity and documentation standards your experimental models require. All compounds are supplied for research use only and are not intended for human, veterinary, or therapeutic application.
Frequently asked questions
What challenges make peptide hormone research models difficult to develop?
Peptide hormones undergo complex precursor processing, diverse post-translational modifications, and rapid post-collection degradation, all of which complicate accurate detection and functional modeling. Processing and PTM complexity00063-5) means that even well-designed assays can measure the wrong molecular species if sample integrity is not controlled.
How does mass spectrometry peptidomics address peptide hormone discovery?
It enables high-throughput, unbiased detection of peptides and their modifications across complex biological matrices, but requires immediate sample inactivation to avoid confounding results. Without heat inactivation, over 95% of detected peptides00063-5) may represent degradation fragments rather than native bioactive hormones.
Why are conditional receptor knockout models important in peptide hormone research?
They isolate receptor-specific physiological contributions with a precision that pharmacological blockade cannot achieve, particularly for peptides acting across multiple tissues simultaneously. Conditional CNS GLP-1R knockouts demonstrated this directly by clarifying the anatomical basis of peptide-conjugate-induced weight loss.
How do computational models complement experimental peptide hormone research?
Computational cleavage prediction narrows the candidate peptide space efficiently, but every predicted fragment requires pharmacological and in vivo validation before it can be assigned biological relevance. Studies mapping over 2,600 PCSK1/3 fragments illustrate both the power and the validation burden of this approach.
What role does pharmacokinetic modeling play in peptide hormone research?
PK modeling relates systemic peptide exposure to clearance, distribution, and half-life parameters, providing the quantitative framework needed to design preclinical dosing studies and support translational extrapolation. Mechanistic PK models using intravenous data and in vitro clearance assays are currently the most reliable approach for therapeutic peptide candidates.